WHY IS YOUR IMMUNITY NOT SIMPLY A NUTRIENT DEFICIENCY, A STRESS RESPONSE OR JUST A BUG?
IMMUNITY, SINGLE NUCLEOTIDE POLYMORPHISMS AND WHAT THIS MEANS FOR YOUR IMMUNE SYSTEM
What does a SNP mean for you and your health?
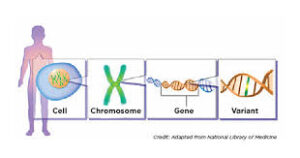
Single nucleotide polymorphisms, frequently called SNPs (pronounced “snips”), are the most common type of genetic variation among people.
Each SNP represents a difference in a single DNA building block, called a nucleotide.
Most commonly, these variations are found in the DNA between genes.
All types of SNPs can have an observable phenotype or can result in disease:
SNPs in non-coding regions can manifest as a higher risk of cancer.
They may also negatively affect mRNA structure and disease susceptibility.
The H4 receptor is a histamine receptor that is predominantly activated in blood cells.
Histamine is a ubiquitous messenger molecule released from mast cells, (enterochromaffin like cells), and neurons and is involved in immune responses.
Additionally, it regulates gut function and is a neurotransmitter that communicates messages for the brain, spinal cord and the uterus.
-
Histamine causes contraction of smooth muscle, especially the lungs, uterus and stomach.
-
It also activates dilation of blood vessels, lowers blood pressure and stimulates the secretion of gastric acid (HCL) in the gut.
Its various actions are mediated by a family of histamine receptors that are a subset of the G protein-coupled receptor superfamily.
Common symptoms of histamine intolerance
-
headaches or migraines
-
nasal congestion or sinus issues
-
fatigue
-
hives
-
digestive issues
-
irregular menstrual cycle
-
nausea
-
vomiting
The protein is thought to play a role in inflammation and allergy and immune.
Their activation is regulated by stimuli such as IFN, TNF-alpha, and IL-6.
Activation of this receptor mediates chemotaxis of mast cells and eosinophils.
In addition, it controls cytokine release from dendritic and T-cells.
Recall from our previous posts the role that T-Cells play in active immunity and the control of cytokine storms.
MTHFR A1298C & MTHFR 677T
The common effects of MTHFR are widely researched and reported.
SNPs in these genes result in deficiencies in methylation, which is the process that allows your body to assimilate your vitamins and minerals efficiently.
Additionally, they impair production of glutathione (our body’s master anti-oxidant).
Predominantly, these polymorphisms DISRUPT THE FOLATE CYCLE which negatively affects the homocysteine cycle as well as many other metabolic pathways.
This causes an inability for our body to utilise B12 efficiently, reduces the ability to detoxify harmful substances like toxins, drugs, alcohol and environmental toxicants.
MTHFR mutations are linked to an array of conditions.
Common findings are Cancer, Cardiovascular Disease, Thrombosis, Stroke, adverse pregnancy outcomes such as pre-eclampsia, still birth & neural tube defects.
Also noted are other Neurological disorders such as depression & anxiety.
Dr. Andrew Rostenberg DC
“For reasons we are still trying to understand, viral DNA becomes permanently part of our own DNA. This process is called horizontal or lateral gene transfer when one organism’s DNA is transferred into the DNA of a separate organism. Because this horizontal gene transfer is so common, scientists have determined that about 40% of our DNA doesn’t actually belong to us. There is so much residue from viruses (and other microorganisms too) that it occupies a large part of our genetic code. It is from this point of view that we can now start to look at the big issue with methylation problems and chronic viral infections.”
It is a commonly documented fact that stress negatively alters our immune response, coupled with the fact that a deficiency in methyl groups alters the epigenetics within our cells; Methylation can change the activity of a DNA segment without changing the sequence. (Adding a methyl group tightly wraps DNA, thus turning genes off. Removing the methyl group loosens the wrapping and enables genes to be expressed if they are not otherwise deactivated).
Dr. Andrew Rostenberg DC
“FUT2 genetic issues deplete the body of L-lysine. Without L-lysine we cannot use our body’s supply of B6 and that creates additional biochemical problems. In short, we need Vitamin B6 and L-Lysine to kill viruses so if you are fighting a chronic virus, make sure you support those pathways with additional supplementation.”
Also, individuals with slow MTHFR pathways or those who may be folate deficient due to chronic stress need to make sure they get at least 400-800mcg per day of methyl folate (5-MTHF). More is not always better, and I recommend a lower dose to my patients unless proven otherwise necessary.
We can thus observe how a viral infection can easily replicate in a person who is stressed, experiences mutant polymorphisms (of which there are many, not just MTHFR) and how our immune function has been suppressed by these mechanisms.”
Why are epigenetics & nutrigenomics important to our immunity, health & wellbeing?